Yeast prions
21 October 2014
Part II
Yeast prions
In recent years, studies on prions present in baker’s yeast, the so-called Saccharomyces cerevisiae, also brought plenty of new information.
When examining this problem it is necessary to stress that yeast prions have no relation to mammal prions and do not cause any diseases.
The name corresponds only to their common feature, namely the ability to take two different conformations without changing the amino acid sequence and, in certain circumstances, to accumulate in the form of β-amyloid plaques.
Of course, yeast prions do influence the cell by causing various phenotype changes, but only in yeasts.
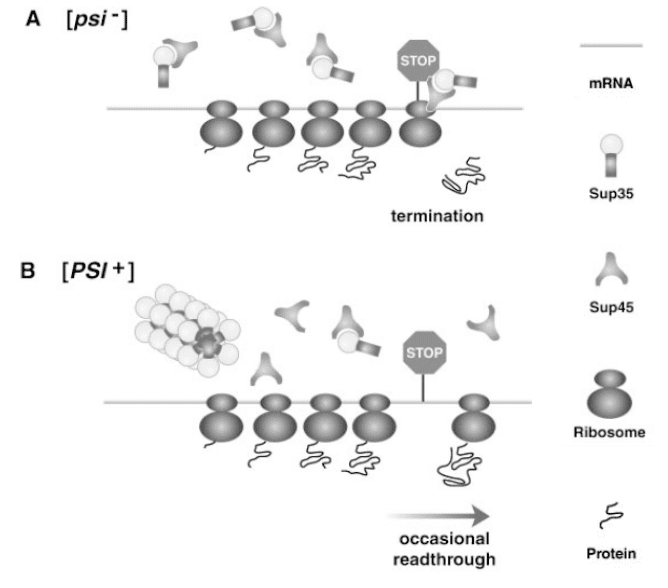
Fig. 7 Scheme of relationship between the Sup35 protein and the [PSI+] agent.
Adapted from Uptain and Lindquist, 2002
Currently, one of the most explored yeast prions is the [PSI+] agent (Cox et al., 1988). Yeast infected with this prion more often omit stop codons located at the terminus of the encoding part of the gene. It is more frequent, therefore, that there are longer mRNA particles in such cells, and at the same time there are longer proteins with additional amino acids at the C-terminus of the polypeptide chain. Intensive studies revealed that the essence of the [PSI+] agent is the Sup35 protein (Wickner, 1994). This protein is an element of an intricate complex revealing the ribosome at a proper time (Stansfield et al., 1995). When the Sup35 protein is unable to function properly, the complex cannot separate the mRNA ribosome at the appropriate moment, thus creating a longer polypeptide chain.
Yeast demonstrating such a feature are marked with the [PSI+] symbol, and the unchanged cells are marked with the [psi-] symbol (Fig. 7).
The already well-known mutation in the ADE2 gene that encodes one of the enzymes necessary to biosynthesise adenine significantly facilitated the [PSI+] agent study.
This mutation introduces the stop codon in the ADE2 gene too early and prevents the creation of a full length enzyme.
Such an enzyme cannot participate in the biosynthesis of adenine and causes accumulation of a semi-product that – luckily for the investigators – has a red colour (Silhankova, 1972).
The scientists conducted an experiment consisting in the insertion of the [PSI+] agent into the cells with the previously specified mutation in the ADE2 gene.
As expected, it turned out that yeast had a normal colour, unlike the [psi-] cells (Cox et al., 1980).
An explanation of this may be the fact that the [PSI+] agent caused an omission of the early sop codon (mutation in the ADE2 gene) and led to the creation of functional particles of the enzyme necessary to biosynthesise adenine – no accumulation of the red pigment was observed (Fig. 8).
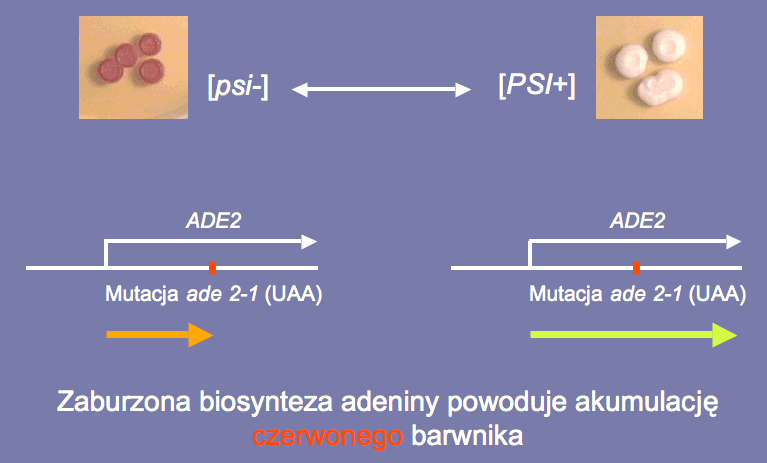
Fig. 8 A comparison of [psi-] and [PSI+] yeast.
The ADE2 gene mutation in [psi-] yeast causes a red pigment to accumulate in it.
The phenotype [PSI+] causing stop codons to be omitted restores the normal appearance of the yeast colony.
The Sup35 protein generally consists of three domains – N, M and C (Fig. 9). The C-domain is the core part of the protein that has an ability to release ribosomes and it was shown that a protein without the N- and M-domains can perform its function in the cell equally well. On the other hand, the N-domain is a part that plays no role in the context of proper protein Sup35 function; however, this very part is responsible for the prion nature of Sup35 protein (Derkatch et al., 1999). This region includes many amino acids with polar ancillary groups (Derkatch et al., 2004), which seems to be characteristic of prion proteins in yeast. This conclusion results from a comparison of several prion proteins in different fungus species e.g. in baker’s yeast.
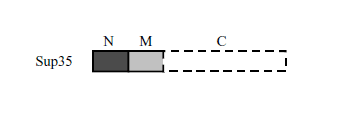
Fig. 9 Sup35 protein domains organisation scheme.
To confirm the role that the N-domain plays in the creation of the prion form of the Sup35 protein, the investigators conducted an experiment aiming to put a chosen protein into the prion state i.e. to change its spatial structure without changing its amino acid sequence (Li and Lindquist, 2000). A variant of glucocorticoids receptor occurring in rats was chosen for this experiment. This receptor is also a constitutively active transcription agent marked as GR526. The above-mentioned protein activates the transcription of genes under the control of the GRE nucleotide sequence.
It is vital to point out that the protein that cannot be found in yeast was deliberately chosen to increase the reliability of the study. The investigators construed an artificial gene, fusing together a fragment of the SUP35 gene encoding the N-domain and M-domain with the GR526 encoding gene. Such an artificial gene produces a protein with N- and M-domains from the Sup35 and GR526 (instead of the C-domain of Sup35 protein). It was expected that if the N-domain was indeed responsible for the conformation change in the protein, the GR526 would lose the function of a transcription agent due to the change of shape.
In order to detect this phenomenon, the investigators created one more construction that consisted in joining the GRE nucleotide sequence with the lacZ gene encoding the β-galactosidase.
The yeasts in which this experimental system was introduced were inoculated into a culture medium, which included a colourless β-galactosidase substrate that turns blue under the influence of the enzyme.
White colonies in the culture would confirm that the N-domain is responsible for the creation of yeast proteins’ prion forms e.g. Sup35 protein (Fig. 10).
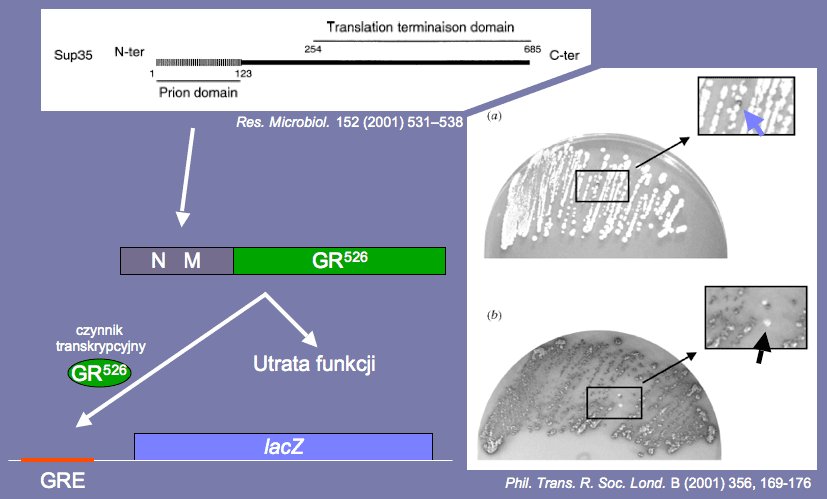
Fig. 10 Artificial protein scheme (N-domain and M-domain from the Sup35 and GR526 proteins), alternative pathways of the protein (loss of functions or binding with the GRE and creation of the β-galactosidase from the lacZ gene), and a photo of plates with spontaneously appearing yeast colonies with different colours.
Indeed, white colonies appeared on the blue background. However, the remaining part of the experiment was interesting. After transplanting the white colonies into a new culture medium, spontaneously appearing blue colonies of yeast were observed. A possible explanation for this is the fact that prions can spontaneously change their conformation state. It seems that proteins from this group are able to change their state from normal to prion without any external stimuli. The most recent studies indicate that the frequency of the spontaneous conformation change in proteins such as Sup35 protein is between 10-5 and 10-7. Of course, some factors i.e. the increase of temperature, addition of the [PSI+] cell lysate, addition of the N-domain and numerous mutations, may increase this frequency (Uptain and Lindquist, 2002). Taking into consideration the influence of the [PSI+] agent, it seems unlikely to bring any benefits to the cells. However, having the [PSI+] prion may be the last and only resort for yeasts with multiple nonsense mutations. In the case of such yeasts, omitting most stop codons (even those localised at a proper place) is better than early stopping of the protein synthesis (True and Lindquist, 2000).
It is a hypothesis that can partially explain the presence of prion [PSI+] in yeasts, although it is rather difficult to imagine a similar explanation for the presence of prions in mammals.
Prion [PSI+] is also significant to phenotypic diversity. The differences between the [psi-] and [PSI+] strains become particularly noticeable in harsh environmental conditions, and the phenotypic diversity caused by the presence of the [PSI+] factor can be considered a stage preceding the increase of genotype diversity.
Specific examples are provided by studies on yeast growth conducted on growth media containing bleomycin, anisomycin or benomyl – these are compounds used in, among others, cancer therapy (True & Lindquist, 2000; True et al., 2004). It turned out that common yeasts were not able to grow on a growth medium containing bleomycin and anisomycin, whereas the cells with the [PSI+] factor were doing very well.
The results were different, however, when benomyl was added to the growth medium. In this case, the [psi-] cells were doing much better than the ones with the [PSI+] factor.
This very well illustrates the nature of this prion. Although its presence allows the cell to survive in some cases, it is difficult to predict what these conditions are, especially considering the fact that in other cases, the [PSI+] factor makes it virtually impossible to survive.
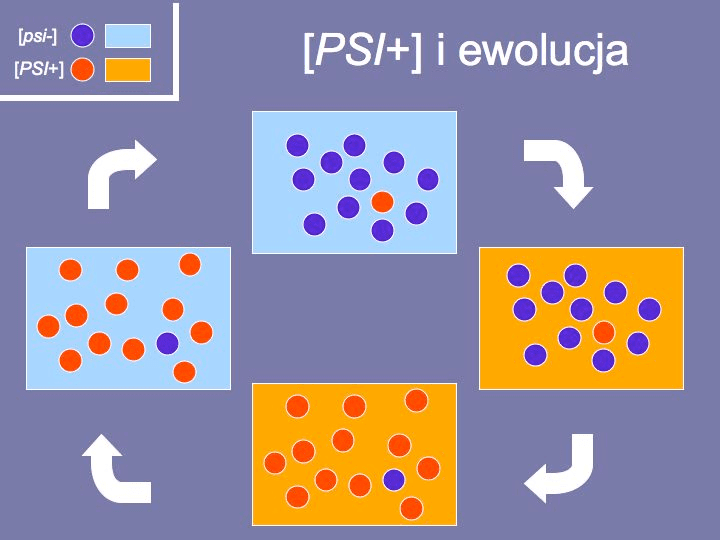
Fig. 11 The significance of the [PSI+] factor in yeast evolution. The dots represent yeast cells and the fields represent the environments. An optimal environment for dark blue yeasts is marked as blue. Red yeasts are best suited for the environment marked as orange. More detailed explanations are provided in the text.
In recent years, a hypothesis has been proposed that assumes the connection between the [PSI+] factor and the yeast genome evolution (True et al., 2004).
As previously mentioned, the spatial structure of the Sup35 protein can spontaneously alter with certain frequency. A normally functioning protein can adopt a conformation characteristic of a prion type. In regular environmental conditions, this change does not benefit the cell in any way.
However, a sudden change of environmental conditions may lead to a situation where the only way to survive is to possess the [PSI+] factor. The [PSI+] cells are favoured in such conditions and start to dominate in the population. As the number of cells increases, the [psi-] cells begin to spontaneously appear. Despite not being favoured in these environmental conditions, if the environmental conditions revert to their previous state, the cells lacking the [PSI+] factor become once again better suited for the surrounding conditions. Obviously, in such a case the [PSI+] cells become the minority (Fig. 11)
The mechanism described above allows the yeast cells to quickly adapt to the current environmental conditions. Unlike genetic changes, the transition from the [psi-] state to the [PSI+] state does not take much time and, if necessary, allows to revert to the previous state. It can definitely be said that this mechanism of adapting the phenotype to the environmental conditions allows the yeast cells to minimise not only the risk but also the costs of evolution.
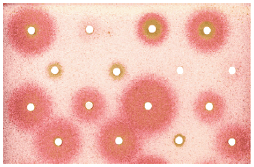
Fig. 12 Yeasts used in studies on anti-prion drugs.
The growth medium surface in a Petri dish becomes covered in full with yeasts carrying a mutation in the ADE2 gene (see Fig. 8) and showing the [PSI+] phenotype. Blotting paper circles are then placed soake with the studied chemical compounds (potential drugs). The circles around which a red colour appears are soaked with the compounds that change the Sup35 protein conformation (losing the ability to skip the stop codon in the ADE2 gene, reverting to the normal structure of the Sup35 protein). These compounds are then examined for the ability to restore the proper PrP protein conformation in mammals. Adapted from Saupe, 2003
The last few years have enabled a synthesis of studies on prions in mammals and yeasts.
For quite some time now, various chemical compounds are being discovered that seem to stop the progression of prion diseases in mammals, but most of all in humans. However, conducting large-scale studies is difficult due to safety reasons. After better understanding the features of yeast prions, the researchers are starting to use yeasts for the initial assessment of medicinal properties of various chemical compounds, approving only the most promising ones for clinical research (Saupe, 2003).
Summary:
Change in phenotype associated with prions is increasingly being considered to be a form of genetic information, although nucleic acids do not take part in its transfer – from this perspective, some researchers qualify it as an epigenetic phenomenon (True et al., 2004), much like a histone code (post-translational modifications of histones that cause various changes, not Fig. 12 Yeasts used in studies on anti-prion drugs. The growth medium surface in a Petri dish becomes covered in full with yeasts carrying a mutation in the ADE2 gene (see Fig. 8) and showing the [PSI+] phenotype.
Blotting paper circles are then placed soaked with the studied chemical compounds (potential drugs). The circles around which a red colour appears are soaked with the compounds that change the Sup35 protein conformation (losing the ability to skip the stop codon in the ADE2 gene, reverting to the normal structure of the Sup35 protein). These compounds are then examined for the ability to restore the proper PrP protein conformation in mammals. Adapted from Saupe, 2003 only in the chromatin state, but also the cellular state; Jenuwein & Allis, 2001). The prion-related problems are being intensively investigated in relation to the diseases that many people and animals of significant economic importance suffer from, as well as for purely scientific reasons. These studies may in fact contribute to discovering the mechanisms of one of the most mysterious phenomena of modern biology. They will certainly allow for expanding the knowledge on the relation between the amino acid sequence and the spatial structure of proteins – an intensively investigated problem that connects many scientific fields.
Author: Takao Ishikawa